By Lambert Strether of Corrente.
This post will do what it says on the tin, and that’s all it will do. Sadly, I actively pursued a state of non-bafflement with genomic software documentation, but after some hours of study, even the rudiments are beyond me. So there will be virtually nothing about genomics in this post (“My eyes clade over.”) I will focus only on the institutions that enable genomic surveillance to be done. I will first allow CDC to define the relevant terms of art. From CDC, “What is Genomic Surveillance?“:
- Mutation: A mutation refers to a single change in a virus’s genome (genetic code). Mutations happen frequently but only sometimes change the characteristics of the virus.
- Lineage: A lineage is a group of closely related viruses with a common ancestor. SARS-CoV-2 has many lineages; all cause COVID-19.
- Variant: A variant is a viral genome (genetic code) that may contain one or more mutations. In some cases, a group of variants with similar genetic changes, such as a lineage or group of lineages, may be designated by public health organizations as a variant of concern (VOC) or a variant of interest (VOI) due to shared attributes and characteristics that may require public health action.
- Genomic Sequencing: Scientists use a process called genomic sequencing to decipher the genetic material found in an organism or virus. Sequences from specimens can be compared to help scientists track the spread of a virus, how it is changing, and how those changes may affect public health.
- Genomic Surveillance: Viruses can be tracked using genomic sequence data collected by CDC and its partners. Effective surveillance does not require the sequencing of a specimen from every COVID-19 case. Instead, scientists rely on collecting enough sequence data from representative populations to detect new variants and monitor trends in circulating variants.
For our purposes (i.e., not pure science) genomic sequencing is what one does to prepare for genomic surveillance. CZ GEN EPI explains further in its Help Center:
To facilitate surveillance efforts, SARS-CoV-2 viruses that are closely related and share signature mutations (genetic changes) are tracked through lineages or variants. A lineage is a group of closely related viruses that evolved from a common ancestor and, thus, share genetic history. A variant refers to a virus with mutations relative to the original SARS-CoV-2 virus detected in 2019. Certain variants with a defining set of mutations can be of more public health importance than others. For this reason, SARS-CoV-2 variants have been named and tracked by Pango, Nextstrain, and GISAID. Each of these platforms has their own nomenclature system that highlights specific virus mutations, but the Pango lineage and Nextstrain clade nomenclatures are the most widely used. When a given variant is demonstrated to be a public health threat, namely ‘variants of concern’ (VOC), it is named following the Greek alphabet (Alpha, Beta, Gamma, Delta, etc). The World Health Organization (WHO) uses this Greek letter nomenclature system to label VOC, which makes it easier to discuss SARS-CoV-2 dynamics and public health responses with general audiences.
So GISAID, Pango, and NextStrain are the most important institutions. I’ll first look at them, in that order, providing a vacuously high-level description of what they do, then pointing to the institutional problems of each. I’ll conclude with a brief rant.
GISAID
From the GISAID About page:
The GISAID Initiative promotes the rapid sharing of data from all influenza viruses and the coronavirus causing COVID-19. This includes genetic sequence and related clinical and epidemiological data associated with human viruses, and geographical as well as species-specific data associated with avian and other animal viruses, to help researchers understand how viruses evolve and spread during epidemics and pandemics.
GISAID does so by overcoming disincentive hurdles and restrictions, which discourage or prevented sharing of virological data prior to formal publication.
The Initiative ensures that open access to data in GISAID is provided free-of-charge to all individuals that agreed to identify themselves and agreed to uphold the GISAID sharing mechanism governed through its Database Access Agreement.
(GISAID stands for Global Initiative on Sharing Avian Influenza Data. Clearly it has moved beyond influenza.)
It’s clear that GISAID has served its archival function very well, from the very beginning of the pandemic:
Today is the 1st anniversary when GISAID learned from China CDC: "It is a novel coronavirus."
36 hrs later, the first genome sequence of the virus was sent to GISAID and released to the world. This data sharing🙏 enabled diagnostic tests and vaccine dev. at unprecedented speed.
— Vaughn Cooper (@vscooper) January 8, 2021
Kudos given, Wikipedia (sorry) describes GISAID’s governance:
GISAID’s administrative affairs are overseen by a board[46] comprising Peter Bogner, and German lawyers Jörg Paura and Christoph Wetzler. Scientific oversight of the initiative comes from its Scientific Advisory Council made up of directors of leading public health laboratories including all six WHO Collaborating Centres for Influenza, and directors of animal health reference laboratories for research on avian influenza for the World Organisation for Animal Health and the Food and Agriculture Organization of the United Nations.
I’ve gotta say, after our horrid experience with WHO and aerosol transmission, that I’m skeptical of any organization that’s WHO-heavy. And a board, any board, with only three people, two of whom are lawyers? I dunno…. But the real issues are governance and access. From The Economist:
[T’his small non-profit organisation is a mighty force in the storage and sharing of genetic data about pathogens…. GISAID has received millions of dollars from the Rockefeller Foundation, a philanthropic organisation; the World Health Organisation (who); and the Coalition for Epidemic Preparedness Innovations, a foundation that funds vaccine research. It has also received donations from pharmaceutical companies. In the first year of the pandemic, the who gave GISAID $1.7m; pharmaceutical firms gave another $1.7m. Donations have continued to roll in, enabling the platform to scale up. By April 2021, 1m coronavirus sequences had been posted to GISAID. In June 2021 the Rockefeller Foundation gave it another $5.1m.
That’s not very much money, in the great scheme of things. More:
Some funders worry about a lack of transparency in the governance of GISAID, especially over the identity of its board members. One funding organisation which asked to remain anonymous describes GISAID as “opaque”. Many, though, understand the organisation to be run mostly by one man: Peter Bogner, its founder. Mr Bogner, a former television-studio executive, is understood to be based in California. (GISAID also has an administrative base in Germany run by a charity, Freunde von GISAID. e.V., or “Friends of GISAID”.)
Nothing sketchy there! (The Economist also says that it’s Big Pharma that’s raising the “transparency” issue, so, er….) And then there’s the question of how open the access really is. Still from the Economist:
On March 21st it emerged that GISAID had revoked the access of a group of international scientists who had been working on Chinese covid data. The argument centred on a dispute over whether they had broken the rules governing use of the database. Their access has since been restored. But the row inspired other scientists to say that they had also had their access to GISAID removed, hampering public-health work.
For example:
Angie Hinrichs, a researcher at the University of California, Santa Cruz, is among those scientists who had her access to GISAID genomic sequences restricted without explanation. Her limited access obliged her to spend 750 hours downloading sequences in tiny chunks during the pandemic, she says.
And:
Bede Constantinides, a senior researcher at the University of Oxford, says that during covid he worked on a system that automated the reporting of lab sequence data. When he asked GISAID if his system could be made to talk to its one—so that data from Britain’s National Health Service could be shared automatically—he received no reply and had his account blocked from uploading to GISAID. GISAID is now “mostly useless” to him, he says, adding that his emails continue to go unanswered. Many scientists say they fear taking their complaints public in case they lose access to the database.
It would be bad if GISAID were undergoing a proces of enshittificiation, like so many other online platforms:
Here is how platforms die: First, they are good to their users; then they abuse their users to make things better for their business customers; finally, they abuse those business customers to claw back all the value for themselves. Then, they die.
It does seem, from the testimony of Hinrichs and Constantinides, that GISAID is abusing its lockin. If so, can and will another platform arise? We shall see.
Nextstrain
Here is how Nextstrain defines itself:
Nextstrain is an open-source project to harness the scientific and public health potential of pathogen genome data. We provide a continually-updated view of publicly available data alongside powerful analytic and visualization tools for use by the community.
Nextstrain provides an open-source toolkit enabling the bioinformatics and visualization you see on this site. Tweak our analyses and create your own using the same tools we do. We aim to empower the wider genomic epidemiology and public health communities.
Here is NextStrain’s workflow, according to a presentation at CDC:
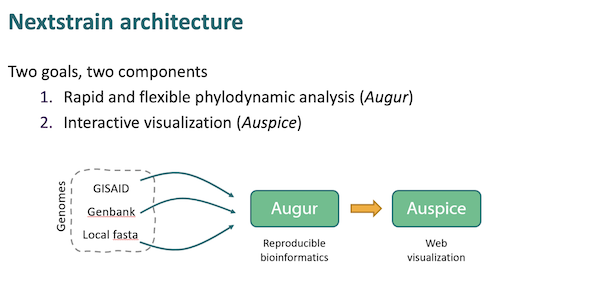
As you can see, the workflow begins at the left a Covid genetic sequence, generally from GISAID. The sequence is then “munged” (technical term) into “reproducible bioinformatics” and displayed to the user. The visualization looks like this:
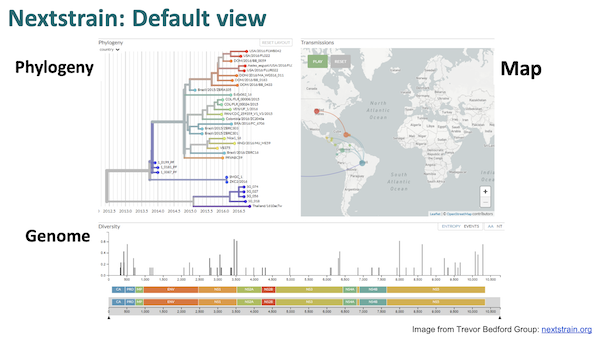
Remember Angie Hinrichs? Here she is again, performing the key role in the “munging”:
Time for a long overdue shoutout to one of the unsung heroes of the pandemic—@AngieSHinrichs, the keeper of the @NextStrain SARS-CoV-2 phylogenetic tree. 1/9 pic.twitter.com/OMz1QaMNMy
— Ryan Hisner (@LongDesertTrain) April 9, 2023
And:
Many sequences are full of errors, some of which are really common, & without these errors being masked by @AngieSHinrichs (& maybe others I don't know about, like @firefoxx66?) , the tree would be riddled with errors and hard to make sense of. 7/9
— Ryan Hisner (@LongDesertTrain) April 9, 2023
So the NextStrain SARS-CoV-2 phylogenetic tree is the editorial product of one person, hopefully never hit by a bus and hopefully never succumbing to Covid brain fog. That, to me, is an institutional weakness.
Pango
Pango is a second open source project, although with an entirely different classificaiton system from NextStrain. BMC Genomics:
The Pango dynamic nomenclature is a popular system for classifying and naming genetically-distinct lineages of SARS-CoV-2, including variants of concern, and is based on the analysis of complete or near-complete virus genomes.
I can’t find a pretty workflow diagram for Pango, but their software page makes the workflow evident:
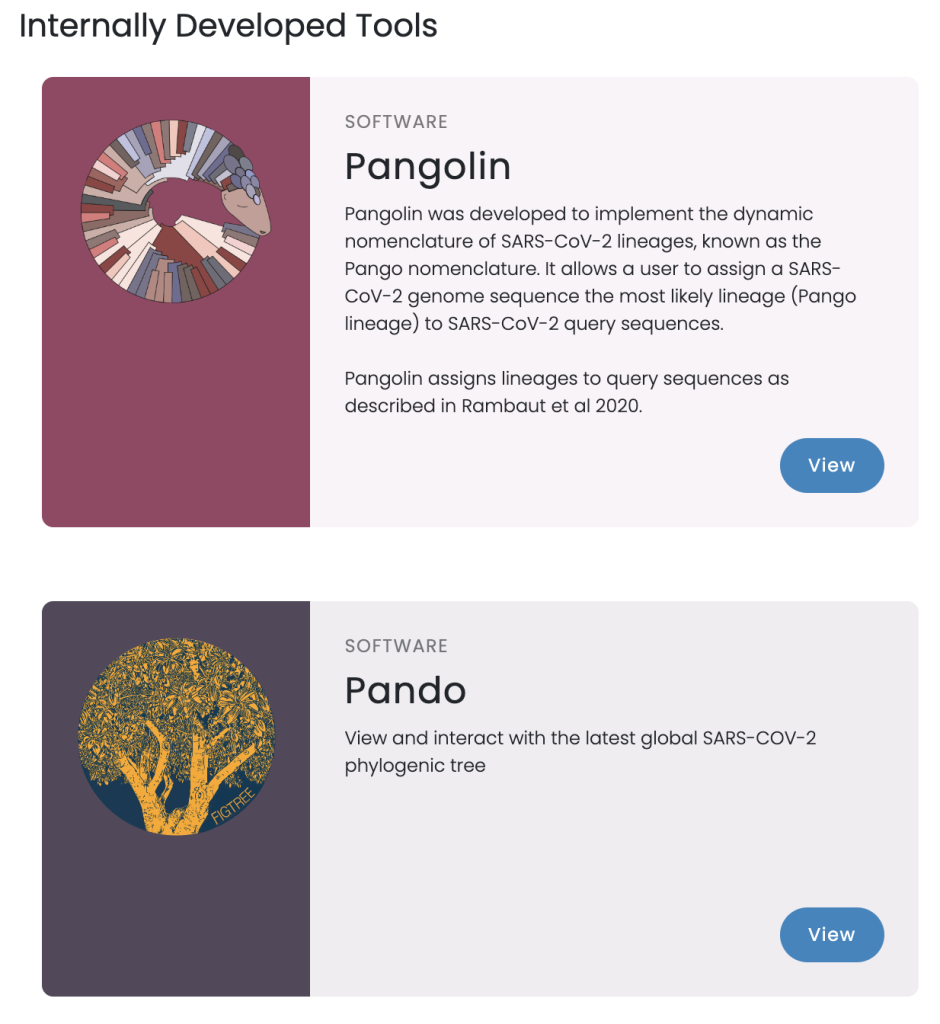
Sequence input from (most likely) GISAID; “munging” in Pangolin; visualization in Pando.
Pango is the system the CDC uses to update its more-or-less weekly variant charts. And Pango has exactly the same institutional weakness as NextStrain. As I wrote back in October 2022:
Now let’s look at the institutional set-up for Pangolin (and please note that I have nothing but the utmost respect for the skills of the developers, or the power and beauty of their work). From MIT Technology Review:
[the Pangolin project is] a GitHub page staffed by a handful of volunteers around the world, led primarily by a PhD student in Scotland.
Those volunteers oversee a system called Pango, which has quietly become essential to global covid research. Its software tools and naming system have now helped scientists worldwide understand and classify nearly 2.5 million samples of the virus.
Researchers, public health officers, and journalists around the world use Pango to understand covid’s evolution. But few realize that the entire endeavor—like much in the new field of covid genomics—is powered by a tiny team of young researchers who have often put their own work on hold to build it.
Many of the foundational tools for tracking covid genomes have been developed and maintained by early-career scientists like O’Toole and Scher over the last year and a half. As the need for worldwide covid collaboration exploded, scientists rushed to support it with ad hoc infrastructure like Pango. Much of that work fell to tech-savvy young researchers in their 20s and 30s. They used informal networks and tools that were open source—meaning they were free to use, and anyone could volunteer to add tweaks and improvements.
“The people on the cutting edge of new technologies tend to be grad students and postdocs,” says Angie Hinrichs, a bioinformatician at UC Santa Cruz who joined the project earlier this year.
So, just to be clear, CDC has outsourced the essential technology for variant detection to volunteers[5]. (And what is the key characteristic of “grad students and postdocs”? They need to move on.) CDC has bet thousands of lives, perhaps tens or hundreds of thousands, on volunteers. Does that sound like a sensible approach to you? Why the heck, again, can’t CDC get them some kinda budget? What happens when the developer gets a better offer? Or moves to another institution? Do people at CDC think that complex open source software is maintained by little elves? Does this sound like operational capacity to you?
No. It very doesn’t.
Conclusion
GISAID’s open access isn’t always open, and in fact they shut down access to two scientist for no good reason I can see. And maybe I can’t see the reason because GISAID’s operations are “opaque.” Of the two essential projects downstream from GISAID, Pango depends on a tiny team of volunteers (!!), and Nextstrain depends on the curation efforts of one person (!!!). Weak, weak, and weak. Dangerous, dangerous, dangerous. What happens if when the genomic sequencing tools go down, and genomic surveillance can’t happen, when a new variant is multiplying geometrically? If when that happens, we can’t afford to lose a week!
So while the PMC moans and wrings its hands because the rentier-servicing labor aristocrats of Silicon Valley won’t be getting free massages or truffle-infused vegan stylings any more, or the political class loses its mind because we can’t send the Azovs in Ukraine enough tanks to break down for parts and sell on the black market, genuine scientists doing the work on which millions of lives depend should look both ways before crossing the street. What a situation. Meanwhile, some brain genius at the Rockefeller foundation misplaced a decimal point. They said a million, I guess because they looked under the couch cushions, but ten million would buy some redundancy. Maybe a hundred millions would buy tech doc dull normals could use, who knows. What’s wrong with these people?


Australian here ( in case context helps anyone).
Thankyou Lambert. You clearly invested a lot of work in a complex subject. And, its refreshing to commence knowing a post will ‘only do what it says on the tin.’ ( Love that expression, which I believe is English)
So, I get that the utility of this genomic ‘whatever’ has an alleged basis in supporting measures to combat ‘covid’. But my eyes glazed over years ago when genomics of the disease were discussed.Why? Well, because I’m old school. And, I also know how medicine and Pharma prefer to hide behind things all shiny and expensive like genomics, also being specialist fields way behind the access of almost everyones intellect. I’m old school. Show me what I can view under a microscope. Then we can learn about how it works, and how to deal with it. Sound fair? So, when governments began passing Covid mandate legislation, as England did as early as March 2020 IIRC. I naturally thought well everyone relies on source principles don’t they! So, all this important urgent legislation comes about because the doctors saw sick people. They took specimens, they studied those under a microscope, observed behaviour and made hypothesis. Consulted with experts. And so on, up the food chain, until the politicians ‘the last to know’ made a decision about the needs of the community. Fair enough? ALL these decisions, including lockdowns, masks,vaccines and their application and utility – ultimately derive from someone studying the virus. Don’t bother me with genomics. Specimens were taken and collected in a petri. I’ve seen what scores of those look like, for known and common viruses or bacterial infections. What does covid look like under a microscope? Hmm. Can’t find it.
Fast forward 12-18 months. I have in my possession 20-30 freedom of information responses to a question something along the lines of ‘Data on the isolated specimen of covid taken from an infected human in 2019 or 2020- who,what,where,how’
And, where can I view it. These FOI responses came from reputable, federal Health or Medicine Department across many english speaking countries. Australia,US,England, NZ,Canada, Ireland, South Africa, Scotland and others. Every single response was identical and reverted to the administrative default. ‘We have done a thorough search and spoken to the relevant people. We must conclude that no such documents exist’.
I don’t care about genomics and new science blazing a path to glory. I’m just asking the question. Why can’t anyone provide me a tissue sample taken from a sick person? Why?
Further to my penultimate comment. Genomics is a map.It is not the terrain. It is ultimately merely a model on a computer screen. Is it representative of objective reality? Well thats an interesting question but lets agree the ‘jury is out’. A model is way more convenient and useful should someone wish to have a vested interest and a desire to control. (Thankyou Cassandra for your comment.) A model provides exponential, arguably infinite opportunity to fashion narratives or manage data that interferes with a pre selected narrative. Keep in mind, the only way a genomic ‘model’ of covid can be accurately constructed to function as a map, is by starting with a tissue sample.
The terrain! How is anyone supposed to construct a map without the raw materials?
By way of analogy. If you are in a court room and the prosecution argues they have finger print evidence! Don’t take their word for it. They are making a claim, they must demonstrate everything they rely upon to substantiate that claim. Demand every data point in the trail, commencing with the initial sample, used to determine the final allegation ‘we have evidence’. Fingerprint technology /reconstruction has a tremendous amount of voodoo.
Similarly: if someone screams they have a genomic model – demand the entire chain of evidence. Where’s my tissue sample?
And Christine Massey continues to collect reams of FOIA responses from institutions all over the world. All have said they do not have / did not find the Covid virus. It’s hard to do genomics on nothing.
https://christinemasseyfois.substack.com/?utm_source=substack&utm_medium=web&utm_campaign=reader2&utm_source=%2Fsearch%2Fchristine%2520massey&utm_medium=reader2
There is a lot of good literature out there about this circumstance, but as I do not want to be cancelled off NC (which I have been reading devotedly since the beginning), I will not add more links.
Ask and you shall receive.
https://www.niaid.nih.gov/news-events/novel-coronavirus-sarscov2-images
From 2020. Many more on the linked flickr page.
GISAID lockouts look to me like the actions of an administrator managing bandwidth limitations. Both the users mentioned have what looks like highly automated and heavy throughput pipelines, likely to hammer an access interface if it isn’t built for it. Given the described GISAID background, it wouldn’t surprise me if they don’t have a well developed API, because it would never be a priority of the specific mostly-volunteers involved.
There are commenters who are much more experienced than me that can comment, but as one of the many grad students involved in genomics, this is what I’ve learnt so far:
On the genomics toolkit front, almost all genomics toolkits are the product of individuals or small groups of grad students and postdocs. It’s not just the “cutting edge”. The difference is that some of the more heavily used and longer-lasting products are managed by now-tenured profs in the open source movement.
The highly monopolized and centralized commercial software for genomics work (2-3 companies globally), from what I’ve used, is based on building a nice gui around the same open source backend.
Genomics tools fall out of maintenance, but there’s always new grad students and postdocs building “slightly better” tools for the same tasks, so we’ve survived so far by moving on to whatever is currently being maintained and isn’t obviously broken. Scientific software is mostly a wasteland of orphaned projects, as you can imagine.
> GISAID lockouts look to me like the actions of an administrator managing bandwidth limitations. Both the users mentioned have what looks like highly automated and heavy throughput pipelines, likely to hammer an access interface if it isn’t built for it
Good argument which, however, does not seem to be explained anywhere (that I have found).
Perhaps the lack of explanation/public justification to the rate-limited users, is part of the lack of transparency The Economist complains of.
Totally agree – that lack of transparency and the heavy handed approach was the other aspect that screamed “ornery sysadmin” to me.
It’s almost like this is a national good that should be publicly funded. The horrors
Thank you for this Lambert. Thoroughly (1) unsurprising and (2) depressing. Maybe the root cause is a bureaucratic disincentive to fund ‘new’ things. Who knows. It is certainly non-responsive to mission.
>And what is the key characteristic of “grad students and postdocs”?
It has been a very long time since I moved in those circles, but in my experience, grad students and postdocs do research. Principal investigators write grant proposals and deal with the associated politics. I doubt much has changed.
The problem is that continuity in research, software development, etc, requires a reliable source of ongoing funds that is not tied to one person’s skills in grantsmanship. But generally speaking, the entities with the money do not want to lose their control of the research. And in this case, it seems those entities do not really want to encourage free-range investigation into the covid virus, and they *really* do want to control access to the information that such investigation may uncover.
Can I don my tinfoil hat? The government agencies and “private” organizations that manage, dictate really, funding for research and the businesses, which use the various research tools for profit along with the national and international government health agencies; they are all interconnected, often across state lines, managing vast amounts of resources, not just money, with often the quiet backing of various security agencies.
Covid is an embarrassing, yet very profitable opportunity for wealth and power, for many, but can still lead to not so figuratively hangings and heads on pikes, if it all comes out. Perhaps at a real fall of empire(s) level of unrest.
You say that they’re problems with getting enough Covid funding? How curious.
It still amazes me that people were not satisfied with merely adequate profits or of funding and maintaining research with the right amount of biological security. Nope, they felt like risking civilizational, or at least state, survival for that scrapping up of the last bit the bottom of the barrel.
.
> if it all comes out
Or else it is all out, right here, in plain sight, which is why it is so hard to see. Follow the money, of course, but also follow where there is no money.
> Scientific software is mostly a wasteland of orphaned projects, as you can imagine
Preventing a pandemic with potentially abandoned software projects seems suboptimal from an operational standpoint. We don’t do the military that way. Or banking.
However, if your policy goal is mass infection followed by profitable treatment with pharmaceuticals, this approach fits right in. Population reduction, with a view to reducing the threat to the carrying capacity of the earth, would be a happy side effect (a global application of Rule #2). I hate to think this way, too, but it’s becoming increasingly hard to avoid. Frankly, I’d be happy to be talked off the ledge. Readers?
Thinking it through, there’s no state-funded global competition for pandemic surveillance software, either. Which would imply that the global capitalist ruling class*, and I very much include China, has made up its mind on the matter, as have the variously configured global PMCs** governing classes. TINA.
* Having, themselves, tooled up for what is to come, a la ventilation at Davos.
** They all went to the same schools.
Thought I’d get a laugh from “truffle-infused vegan stylings.” Tough crowd!
For what it’s worth, in Europe the Covid-19 “open data” stuff is concentrated on Covid-19 Data Portal run basically by European Molecular Biology Laboratory (which, regardless of it’s name is also an umbrella organization for 6 actual EMBL laboratories, 6 local alliance institutes and 9 partnership institutes).
The Data Portal does automatic phylogeny for uploaded data as described in Szarvas et al. (2020) using KMA Clausen et al. (2018).